Biomedical engineers have built simple machines out of DNA, consisting of arrays whose units switch reversibly between two different shapes.
The arrays’ inventors say they could be harnessed to make nanotech sensors or amplifiers. Potentially, they could be combined to form logic gates, the parts of a molecular computer.
The arrays’ properties are described in Science.
The DNA machines can relay discrete bits of information through space or amplify a signal, says senior author Yonggang Ke, PhD, an assistant professor in the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory.
“In the field of DNA-based computing, the DNA contains the information, but the molecules are floating around in solution,” Ke says. “What’s new here is that we are linking the parts together in a physical machine.”
Similarly, several laboratories have already made nanotech machines such as tweezers and walkers out of DNA. Ke says his team’s work with DNA arrays sheds light on how to build structures with more complex, dynamic behaviors.
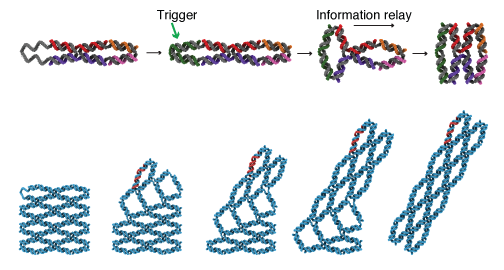
The arrays’ structures look like accordion-style retractable security gates. Extending or contracting one unit pushes nearby units to change shape as well, working like a domino cascade whose tiles are connected.
The arrays’ units get their stability from the energy gained when DNA double helices stack up. To be stable, the units’ four segments can align as pairs side by side in two different orientations. By leaving out one strand of the DNA at the edge of an array, the engineers create an external trigger. When that strand is added, it squeezes the edge unit into changing shape (see illustration).
To visualize the DNA arrays, the engineers used atomic force microscopy. They built rectangular 11x4 and 11x7 arrays, added trigger strands and could observe the cascade propagate from the corner unit to the rest of the array. The arrays’ cascades can be stopped or resumed at selected locations by designing break points into the arrays. The units’ shape conversions are modulated by temperature or chemical denaturants.
For reference, the rectangular arrays are around 50 nanometers wide and a few hundred nanometers long – slightly smaller than a HIV or influenza virion.
To build the DNA array structures, the engineers used both origami (folding one long “scaffold” strand with hundreds of “staple” strands) and modular brick approaches. Both types of arrays self-assemble through DNA strands finding their complimentary strands in solution. The origami approach led to more stable structures in conditions of elevated temperature or denaturant.
In the Science paper, the engineers showed that they could build rectangles and tubes of array units. They also include a cuboid that has three basic conformations, more than the two-dimensional array units with two conformations. Ke says his team is working on larger, more complex machines with three-dimensional shapes, which can be made using the same basic design principles.
The laboratory of Chengde Mao, PhD in Purdue University’s Department of Chemistry contributed to the paper. The co-first authors of the paper are postdoctoral fellow Jie Song, PhD, now at Shanghai Jiaotong University, postdoctoral fellow Pengfei Wang, PhD, and Purdue graduate student Zhe Li.
The research was supported by the National Science Foundation (CAREER DMR–1654485, CMMI–1437301), the Marcus Foundation, the Office of Naval Research (N00014-15-1-2707) and the National Natural Scientific Foundation of China (21605102).