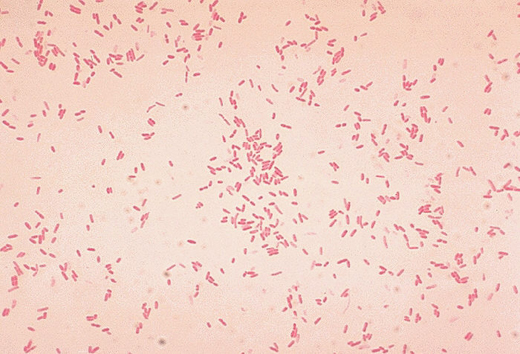
The common waterborne bacterium Aeromonas hydrophila has increasingly been implicated in serious human infections. By correlating clinical and experimental findings with genome sequencing data, scientists have found key factors that distinguish bacteria that can cause necrotizing skin infections ("flesh-eating bacteria") from other bacteria commonly found in freshwater sources.
Joshua Shak, an MD/PhD student at Emory’s Rollins School of Public Health, collaborated with a team of scientists from the FDA, the University of Texas, the University of Maryland, the Maryland VA, and Emory University School of Medicine to examine bacteria of the species A. hydrophila. Together, the group used whole-genome sequencing of clinical Aeromonas strains, followed by corresponding laboratory analysis, to identify disease-causing characteristics.
The complete study is published in the April 23, 2013 edition of the Open Access journal mBio.
The newly published study was a follow-up to a case report titled, "Aminoglycoside-Resistant Aeromonas hydrophila as Part of a Polymicrobial Infection following a Traumatic Fall into Freshwater," published in the March 2011 edition of the Journal of Clinical Microbiology.
For the new study, scientists sequenced the entire genomes of bacteria isolated from an individual infected after falling into Georgia’s Chattahoochee River. The goal was to determine key differences between the bacteria that caused severe disease and bacteria of the same species that are harmless. Findings indicated a number of genes unique to each strain, but one particular strain, identified as E1, contained certain genes that may explain why it was so aggressive and persistent in human infection.
“We did a comprehensive analysis of which genes were in each strain and found a number of genes in the E1 strain that could enhance the ability of this bacterium to cause disease,” explains Shak. “We then performed a variety of laboratory experiments to ensure that what we were seeing at the genetic level was also present at the phenotypic, or behavioral, level.”
Colleagues at the University of Texas Medical Branch analyzed a number of behaviors in the laboratory such as bacterial motility, or the ability to move using whip-like appendages called flagella. Results indicated that the E1 strain was a more dominant swimmer (moving through a semi-solid media) and swarmer (spreading across the surface of media).
Based on the findings, the researchers concluded that the E1 strain was more pathogenic on three different levels:
Clinical: This bacterial strain caused an aggressive, rapidly advancing infection in a human.
Genomic: Examination of the entire genetic sequence revealed genes encoding for toxins, capsules that surround and protect the bacteria, and motility-enabling flagella.
Experimental: When tested in the lab, the E1 strain displayed a superior ability to break open red blood cells, secrete toxins, survive in human blood, move rapidly, and kill mice.
“These findings are fascinating because, although A. hydrophila is common in freshwater and usually harmless, a small subset is more prone to causing disease,” concludes Shak. “Furthermore, our findings highlight the diagnostic potential of whole-genome sequencing. With future advances in rapid sequencing and genome analysis, we will be able to provide timely information on the identity and the disease-causing potential of clinically-isolated microscopic organisms.”
###