Imagine reading an entire book, but then realizing that your glasses did not allow you to distinguish "g" from "q." What details did you miss?
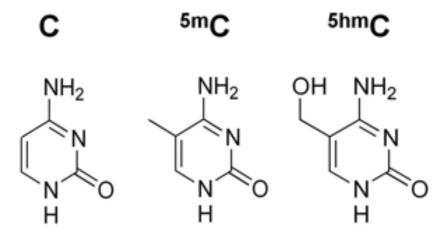
Geneticists faced a similar problem with the recent discovery of a "sixth nucleotide" in the DNA alphabet. Two modifications of cytosine, one of the four bases that make up DNA, look almost the same but mean different things. But scientists lacked a way of reading DNA, letter by letter, and detecting precisely where these modifications are found in particular tissues or cell types.
Now, a team of scientists from the University of Chicago, the Ludwig Institute for Cancer Research, University of California, San Diego and Emory University School of Medicine has developed and tested a technique to accomplish this task. The results were published May 17 in the online edition of the journal Cell.
The team used the technique to map 5-methylcytosine (5-mC) and 5-hydroxymethylcytosine (5-hmC) in DNA from human and mouse embryonic stem cells, revealing new information about their patterns of distribution. These studies have revealed that these DNA modifications play major roles in fundamental life processes such as cell differentiation, cancer and brain function.
"They regulate gene expression and have a broad impact on stem cell development, various human diseases such as cancer, and potentially on neurodegenerative disease. They may even shape the development of the human brain," says Chuan He, professor in chemistry at the University of Chicago.
Scientists have been examining the patterns of 5-mC for decades, as part of the field of epigenetics: the study of the information that lies "on top" of the DNA sequence. However, researchers only recognized that 5-hmC was present at significant levels in our DNA a few years ago. 5-mC is generally found on genes that are turned off and helps silence genes that aren’t supposed to be turned on. In contrast, 5-hmC appears to be enriched on active genes, especially in brain cells. Also, defects in the Tet enzymes that convert 5-mC into 5-hmC can drive leukemia formation, hinting that changes in 5-hmC are important in cancer.
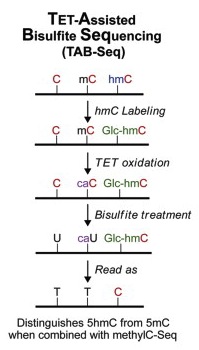
The Cell paper describes a method called TAB-Seq that directly measures 5-hmC, and presents the first map of the entire genome of 5-hmC at single-base resolution. Chuan He and three of his students conceived and developed the technique at the University of Chicago. The other two laboratories of the team, Bing Ren’s Ludwig Institute for Cancer Research/UCSD group and Peng Jin’s group at Emory, applied TAB-Seq to human embryonic stem cells and mouse embryonic stem cells respectively. Jin is an associate professor of human genetics at Emory University School of Medicine.
Previous studies had shown that 5-hmC was found on genes that are turned on. Now, the additional resolution and subsequent research on mouse and human embryonic stem cells reveals that it is found more on the stretches of DNA that control a gene’s activity, rather than the parts of genes that are actually read out into RNA.
"We learned using this new technique that this modification is most abundant in the areas of the genome known as enhancers, which regulate the expression of genes. This potential regulatory role of hmC may explain its importance in embryonic stem cells, and why its disruption may result in the development of leukemia," says Gary Hon, a postdoctoral fellow in Bing Ren's laboratory, who carried out genome-wide analysis of 5-hmC in human embryonic stem cells.
Another difference with 5-mC is that 5-hmC is usually on only one side of the DNA. In contrast, 5-mC is most often found symmetrically. In ES cells overall, 5-hmC is around 14 times less abundant than 5-mC. Even at sites where 5-hmC is the most abundant, it is still present at around one fifth the frequency as 5-mC, the team found using the new technique.
Previous research has found that 5-hmC is 10 times more abundant in brain than in stem cells, so it may have an especially important role there. Jin’s laboratory is using the new technique to finely map 5-hmC in the developing brain.
"To really see the kinds of functions 5-hmC can have, we need to look at how it appears and disappears over time, during processes like brain development. This technique will allow us, and other investigators, to dive in and get that information at high resolution," Jin says.
A patent is pending on the team's invention. The University of Chicago is working with Chicago-based Wisegene to further develop the technology.
An alternative technique for 5-hmC sequencing was described recently in the journal Science.
The research was supported by the National Institute of Health (5R01GM071440-07, 5R01NS051630-07, 5P50AG025688-08, and 5U01ES017166-04), Chicago Biomedical Consortium, Searle Funds at the Chicago Community Trust, Emory Genetics Fund, Simons Foundation Autism Research Initiative, Autism Speaks, and the Ludwig Institute for Cancer Research.
Reference: M. Yu et al. Base-Resolution Analysis of 5-Hydroxymethylcytosine in the Mammalian Genome. Cell (2012) doi:10.1016/j.cell.2012.04.027
