Zoonotic Diseases
How Pathogens Jump Species
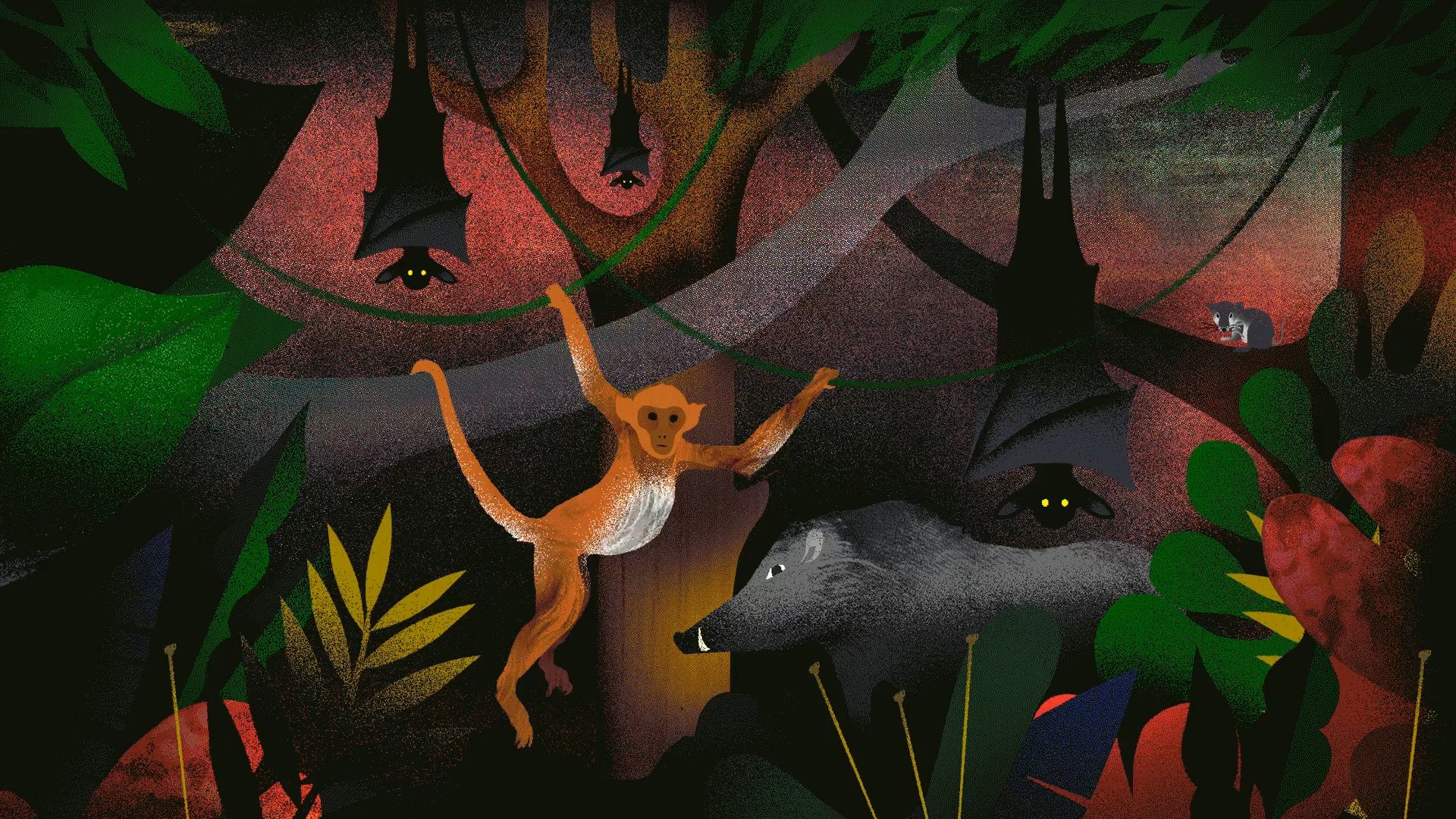

IT STARTS WITH AN ANIMAL. Not a specific kind of animal, and generally not one that is trying to do any harm. But inside that animal lies a pathogen—a microscopic parasite that has spent lifetimes reproducing inside its animal carrier—primed by evolution and ready to take on a new host, should the opportunity arise.

With the growing human population, cities and towns popping up in previously untouched areas, farms and livestock encroaching on wildlife habitats, and rapid climate change, these opportunities abound. The good news is that zoonotic pathogens are making the leap from animals to humans under the spotlight of unprecedented scientific observation.
“We can’t afford to just focus on one pathogen or one animal. It’s really important to get a general understanding of the interactions of different species and how changes in the environment are driving zoonotic disease transmission,” says Thomas Gillespie, a disease ecologist in Emory's Department of Environmental Sciences and the Rollins School of Public Health. "The majority of emerging infectious diseases are coming from wildlife and most of that wildlife is in tropical forests."
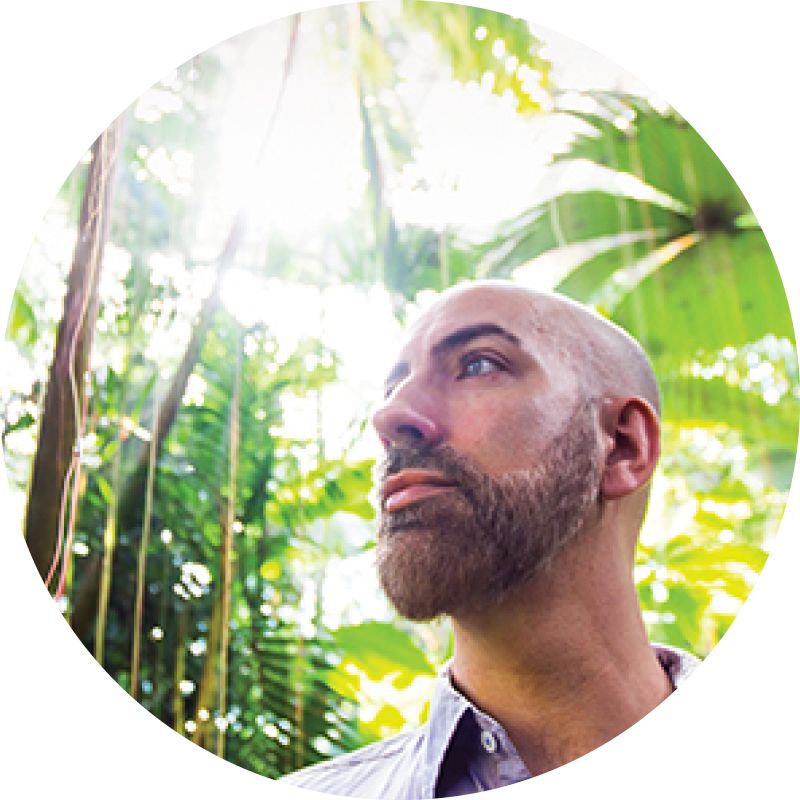

Thomas Gillespie is a disease ecologist in Emory’s Department of Environmental Sciences and an associate professor of environmental health at the Rollins School of Public Health.
Some scientists seek to predict the future of outbreaks with sophisticated mathematical tools and programming. Others go directly to outbreak sites to track, and attempt to contain, the spread of the disease. Still others focus on the mechanisms that allow the pathogens to make jumps between seemingly disparate species. All seek to protect humans from the next unknown threat, whenever and wherever it inevitably emerges.
This emergence is a byproduct of contact with infected animals as well as the nature of viral evolution. Most viruses are programmed for survival within a certain host. This includes factors like the virus’ ability to attach to host cells. Like a microscopic lock-and-key mechanism, proteins on the surface of virus particles are shaped in specific ways that allow them to bind with receptors on host cells.
These interactions can be very specific—they determine which cells are bound (infections of cells in the respiratory tract may cause a cough, while those of the stomach can cause nausea and diarrhea) and which organisms the virus can infect.
If that key changes shape because the virus has mutated, it may be able to open new locks, infect a new host, and cause an outbreak. “How an emerging pathogen spreads through a species tends to be ‘a black box’ until it causes an outbreak among people,” Gillespie says. The Zika virus, for instance, was first identified in monkeys in Uganda in 1947 but was not widely studied until recently, after it started sweeping through human populations.

Thomas Gillespie is a disease ecologist in Emory’s Department of Environmental Sciences and an associate professor of environmental health at the Rollins School of Public Health.
Thomas Gillespie is a disease ecologist in Emory’s Department of Environmental Sciences and an associate professor of environmental health at the Rollins School of Public Health.
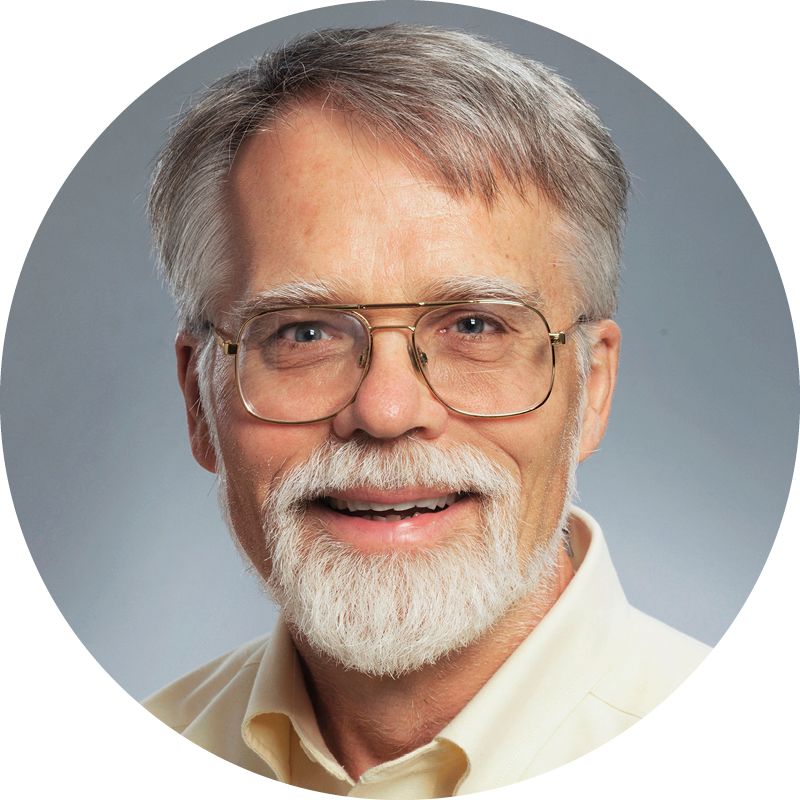

Larry Anderson is a professor of pediatric infectious disease, a member of Emory’s Vaccine Research Center, and Marcus Chair of Infectious Diseases at Emory.
Larry Anderson is a professor of pediatric infectious disease, a member of Emory’s Vaccine Research Center, and Marcus Chair of Infectious Diseases at Emory.
Previous Coronaviruses
COVID-19 is not the first deadly coronavirus to make the jump from animals to humans. Both severe acute respiratory syndrome (SARS) and Middle East respiratory syndrome (MERS) are coronaviruses classified as zoonotic viral diseases, meaning the first humans infected acquired these viruses directly from animals.
SARS was initially present in an as-yet unknown animal reservoir, perhaps bats, and was passed to civet cats, relatives of the mongoose. Evidence shows that as it circulated in the civet cats, it gained mutations that allowed it to cross over into humans in 2002-03. More than 8,000 people worldwide became sick with SARS and 774 died.

Larry Anderson is a professor of pediatric infectious disease, a member of Emory’s Vaccine Research Center, and Marcus Chair of Infectious Diseases at Emory.
Larry Anderson, Marcus Chair of Infectious Diseases at Emory and a member of Emory's vaccine research center, was with the Respiratory and Enteric Viruses branch of the CDC at the time of the 2002 SARS outbreak. Like many new diseases, he says, “SARS started out as a disease of unknown etiology.”
After a doctor identifies a new disease, scientists must figure out the causative agent. From there, it’s a matter of controlling spread. The ease of this control depends on the severity of disease. In the case of pathogens like Ebola, people have severe symptoms. Those exhibiting these symptoms can be quarantined and healthy individuals can avoid contact.
In the case of diseases like the current COVID-19, “transmission happens before serious illness does, which means it’s much harder to use classic public health measures,” says Anderson. Some people who are sick are asymptomatic, so they can spread disease without knowing it.
Evidence suggests that MERS, first identified in 2012 during an outbreak in the Arabian Peninsula, was transferred from bats to camels to humans. Since then, the virus has been endemic in camels of that area and has infected more than 2,000 people and caused more than 700 deaths.

When Viruses Jump Species
Normally, we have specific mechanisms to avoid mutations as our cells reproduce, aptly named cellular proofreading. Viruses, however, are not programmed in this way—they accumulate mutations rapidly and in a variety of ways.
Imagine a genome as an instruction manual, each chapter detailing how to assemble part of a machine. During infection, these chapters tell the cell how to make an entire virus. As the virus replicates inside of a cell, it copies down this manual thousands of times and packages it into new virions which carry those instructions. During the copying process, however, mistakes happen.
Influenza virus, for example, makes approximately one error per new genome—that’s the equivalent to one miscopied word per new instruction manual. In a viral context, that miscopied word can have a big impact and result in the encoded machines severely malfunctioning. A virus can also swap out entire sections of its genome with other viruses of the same type, like swapping entire chapters of the manual. Usually, this is bad for the virus—the new parts don’t fit together and the virus can’t form a productive infection.
Sometimes, however, this works to the virus’ advantage. That’s how the swine flu outbreak occurred in 2009. By swapping sections of its genome, a virus can combine parts of machinery that fit different hosts and subsequently cross species borders, provided that these parts work together successfully.
The mistakes made during replication, whether to a single piece of one protein or in exchange of entire sequences, are random. It’s difficult to know which viruses will evolve to infect new hosts and which species will be responsible for outbreaks.

Fliers and Fast Breeders
Many animal species harbor pathogens— some of which they live with and are largely unaffected by. A common example is bats.
“One quarter of mammal species overall are bats, and each of these myriad bat species carries a suite of different pathogens,” says Gillespie. “Bats are able to host different viruses without getting sick. So bats, and the pathogens that bats carry, are numerous. And bats and humans are both mammals. This relatedness means we’re more likely to get a pathogen from a bat than from a cricket, for instance.”
But bats are far from the only reservoirs of zoonotic pathogens. Influenza virus is housed primarily in aquatic birds, but in 2009 emerged from swine into humans, causing a global pandemic. The “swine flu,” caused by the H1N1 virus strain, which started in pigs killed between 150,000 and 575,000 people worldwide during the first year the virus circulated. Unlike COVID-19, 80 percent of H1N1- virus-related deaths were estimated to have occurred in people younger than 65 years of age—primarily children and young and middle-aged adults.
Rodents are another common host and make particularly good reservoirs, possibly because of their “live fast, die young” strategy.
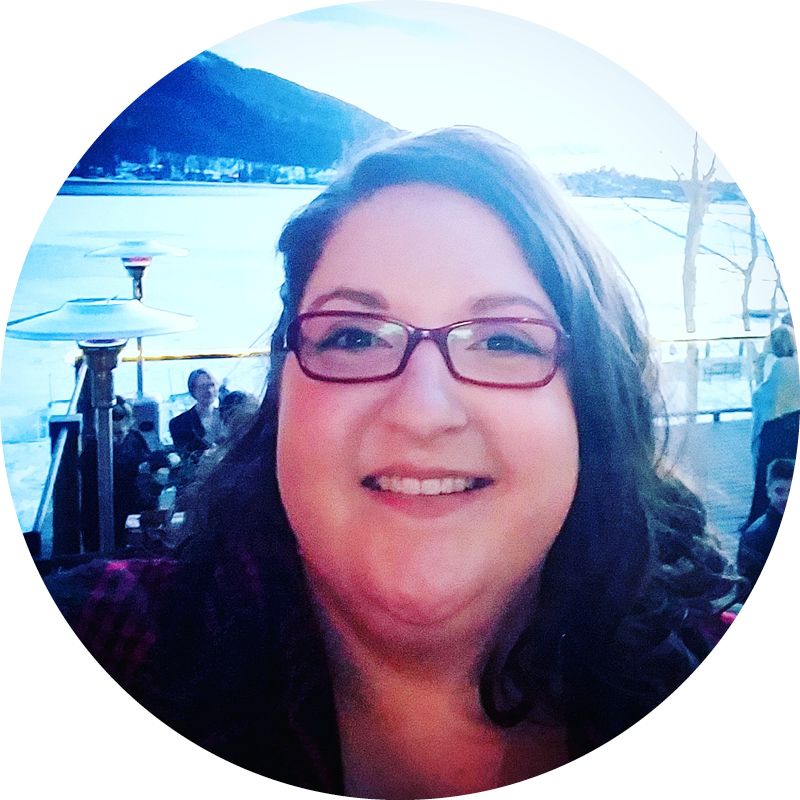

Quantitative ecologist Sarah Bowden is an adjunct professor at Emory and a data scientist contractor with the Division of Global Migration and Quarantine at the Centers for Disease Control and Prevention (CDC).
Quantitative ecologist Sarah Bowden, an adjunct professor at Emory and a data scientist contractor with the Division of Global Migration and Quarantine at the Centers for Disease Control and Prevention, “fast-living species that reproduce quickly and frequently tend to be better reservoirs for pathogens.”
A case in point: Deer mice, which were linked to a hantavirus outbreak in 1993 that caused severe pulmonary disease.
“Previously hantavirus had not been associated with a pulmonary condition,” says infectious disease epidemiologist Robert Breiman, of Emory’s Global Health Institute. “It was a virus that hadn’t been identified before, and it appeared in these deer mice.”
Wild animals that migrate or have large territories also make effective vectors.
“If you cover a broader area, you're inherently more likely to encounter more species you can transmit that pathogen to,” Bowden says.
Zoonosis is a widespread, complex, and multifaceted phenomenon. It does not discriminate between borders, and by nature does not always discriminate between hosts.
Humans, agree scientists, have created conditions for these pathogens to thrive, and rapid human expansion will undoubtedly contribute to the emergence of new diseases.
In some places, it is common to hunt wildlife for food. Rural communities may rely on meat from local forests and savannas—whether chimpanzees, bats, rodents, or other species. These animals can be carriers of diseases, including Ebola, Marburg, HIV/AIDS, and anthrax.
“Anthrax is primarily zoonotic and wouldn’t cause human disease unless there was close contact [between animals and humans],” says Breiman.
Whether an individual is hunting, butchering, or consuming these animals, or is even a child playing near their habitats, this gives pathogens ample opportunity to cross over and wreak havoc.
Public health workers focus on educating and developing safer practices for hunting and butchering.
But domesticated and farm animals can transmit disease as well. Pigs, chickens, turkeys, and waterfowl transmit swine and avian flus. Cows and camels play host to bacterial species like Brucella.
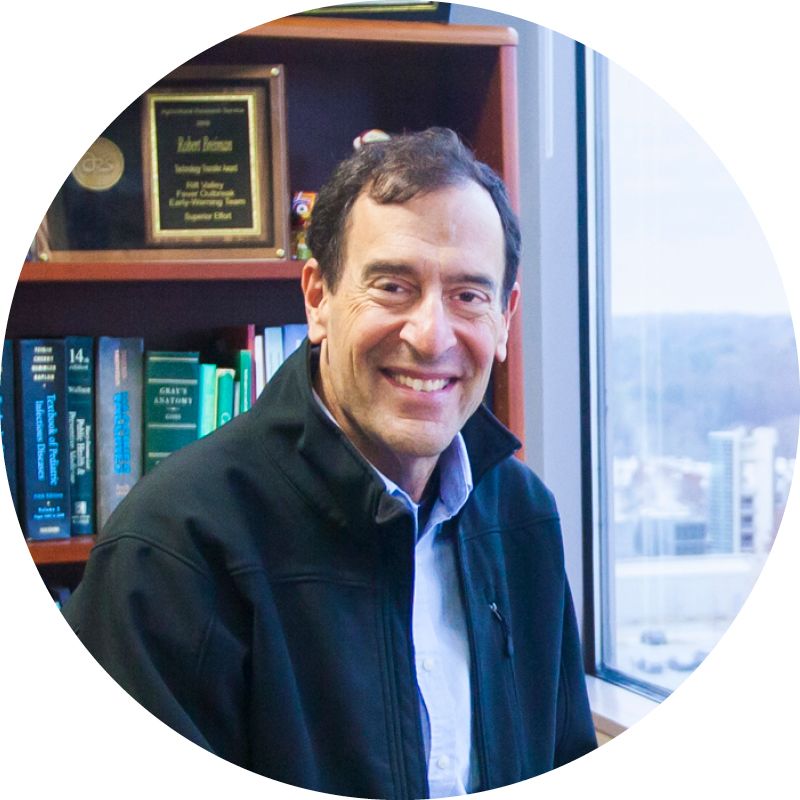

Infectious disease epidemiologist Robert Breiman, of Emory’s Global Health Institute, is also a professor of global health, environmental health, and infectious diseases at Rollins School of Public Health and Emory School of Medicine.
“In parts of East Africa, people really love unpasteurized milk from both cows and camels,” said Breiman, who spent nine years in Kenya as an epidemiologist. “Both animals, if infected with Brucella, can excrete it into the milk,” which can infect those who drink it.
To add to the problem, livestock rarely show symptoms of infection with Brucella so it’s hard to know that the milk poses a risk.
Even if an animal appears sick, that is no guarantee it won’t be consumed. “There will be all sorts of clues that an animal was sick and died, but [people] will go ahead and eat it because it’s nutrition and it costs money,” Breiman says.
In “wet markets” around the world, animals alive and dead, including wild and exotic species, can be purchased for consumption or for use in traditional medicines.
Civet cat, for instance, is a delicacy in some parts of China. “I was told people would eat it in the winter because they felt it increased their immunity to respiratory disease,” Breiman says.
The scales of the pangolin—an armadillo-like animal also sold in “wet markets” that may have played a role in the emergence of the new SARS-CoV-2 coronavirus in humans—are used in medicines. “The meat is thought to have all sorts of benefits,” Breiman says.
Animals in these markets are caged next to each other, which allows them to share pathogens. In the wild, many of these animals would never be close enough for this transfer to occur. The adaptations that happen during infection of a new host can have the unwanted side effect of making them able to infect people, too.
Humans have created an environment in which this is possible, and multiple outbreaks have occurred as a result. The most recent is the 2019 coronavirus.

Connectivity and Climate Change
Sometimes, however, our encounters with animals are unintentional—a result of rapid human expansion and the resultant narrowing of animal habitats.
“We are in an extremely connected world, a world that's vastly more connected than it was even back in the early 2000s when the SARS epidemic happened,” Bowden says. “That is just going to inherently increase the likelihood of new people coming into contact with new things that can make them sick.”
While there are many species known to transmit diseases to humans, there are plenty that we haven’t encountered yet. That is changing as we travel more and expand our reach into untouched areas with remote species and pathogens.
“Things are already there and primed to go, and the storm sort of comes together at the perfect moment,” Bowden says.
But that doesn’t stop researchers from trying to predict outbreak likelihoods from what we do know. Researchers like Bowden have adapted mathematical modeling techniques to better understand where risk of an outbreak lies. “Prediction allows us to look at the relative risk or the likelihood of a pathogen emerging in different areas of the world,” she says.
She looks at different factors—the risk landscape on a spatial scale (where on a map is infection most likely), different known reservoirs (which animals are common carriers, and where are they located), and different pathogens (what germs/diseases are commonly found in an area).
“We can drill through these and look for places where risk is high on all three planes. That allows us to say, ‘Relative to everywhere else in the world, given the large players in a disease outbreak, here’s where we think the risk is highest.’ ”
Many of these factors are changing rapidly, however, and models will have to change to account for that.
Climate change, Bowden says, is “a huge consideration in disease ecology. It is causing species’ geographic ranges to expand or shift. They can bring new pathogens with them or come into contact with ones they hadn’t encountered before. In both cases, we’re likely to experience unforeseen outbreaks.”
Written by Megan Hockman. Illustrations by Brian Stauffer. Design Peta Westmaas. Editor Mary Loftus


Quantitative ecologist Sarah Bowden is an adjunct professor at Emory and a data scientist contractor with the Division of Global Migration and Quarantine at the Centers for Disease Control and Prevention (CDC).
Quantitative ecologist Sarah Bowden is an adjunct professor at Emory and a data scientist contractor with the Division of Global Migration and Quarantine at the Centers for Disease Control and Prevention (CDC).

Infectious disease epidemiologist Robert Breiman, of Emory’s Global Health Institute, is also a professor of global health, environmental health, and infectious diseases at Rollins School of Public Health and Emory School of Medicine.
Infectious disease epidemiologist Robert Breiman, of Emory’s Global Health Institute, is also a professor of global health, environmental health, and infectious diseases at Rollins School of Public Health and Emory School of Medicine.
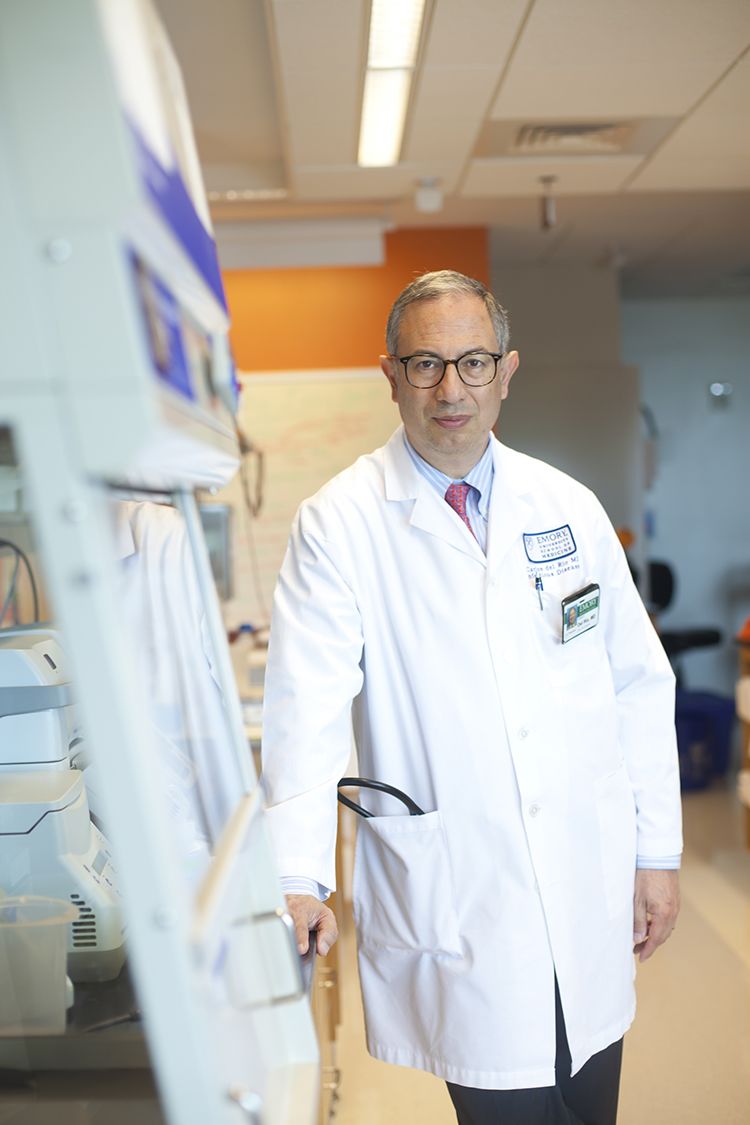
Origin of the Pandemic
“From genetic sequencing data, it appears that there was a single introduction into humans followed by human-to-human spread. This novel virus shares 79.5% of genetic sequence with SARS-CoV and has 96.2% homology to a bat coronavirus. In addition, SARS-CoV-2 shares the same cell entry receptor ACE2, with SARS-CoV. It is still unclear what animal may have served as a bridge for the pathogen to cross from bats to humans. For SARS, evidence shows it was civet cats, for MERS it was camels. While the initial human case of SARS-CoV-2 is yet unknown, early on the Huanan Seafood Wholesale Market was linked epidemiologically.”— JAMA Viewpoint, “2019 Novel Coronavirus—Important Information for Clinicians." Co-authored by Carlos del Rio, MD, Division of Infectious Diseases, Department of Internal Medicine, Emory School of Medicine (at right), and Preeti Malani, MD, MSJ, Division of Infectious Diseases, Department of Internal Medicine, University of Michigan, Ann Arbor; and associate editor, JAMA.



